New Algorithm Identifies Ten Times More Naturally Occurring Antibiotics than All Previous Studies
Drug resistance is a major concern worldwide. Many drugs, including antibiotics of ‘last resort’ such as vancomycin and daptomycin, are Peptidic Natural Products (PNPs) that have an unparalleled track record in pharmacology. Many antimicrobial and anticancer agents are PNPs, but discovery of new PNPs is a difficult challenge – both experimentally and computationally.

In a paper* published in Nature Microbiology on January 22, a team of American and Russian computer scientists described a new algorithm that identified an order of magnitude increase, or roughly 10 times more PNPs than all previous studies. Pavel Pevzner, a professor of computer science and engineering at the University of California San Diego, is the corresponding author on the paper.
PNPs represent one of the last bastions of the complex compounds that, until recently, remained virtually untouched by computational research. These compounds are produced by bacteria and fungi in a battle for survival and they offer great potential to be natural antibiotics. With antimicrobial resistance becoming a global concern and medicine eager for new antibiotics, innovative methods for discovering natural product antibiotics are becoming of utmost importance.
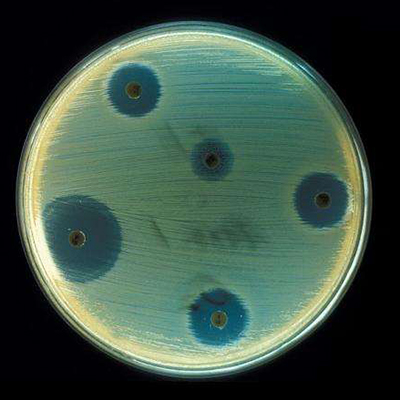
Courtesy: Centers for Disease Control
In the study, researchers describe VarQuest, a novel computational approach for PNP identification. VarQuest can process immense amounts of mass spectrometry data (all spectra of natural products ever generated and made publicly available) in a single run. As a result, VarQuest can be applied in high-throughput discovery pipelines such as the recently launched Global Natural Products Social (GNPS) molecular network. GNPS already contains over a billion mass spectra collected worldwide, a gold mine for future discovery of bioactive compounds. In contrast to existing competitors, VarQuest is able to identify not only known PNPs but also their novel variants, which are sometimes more clinically effective.
VarQuest analysis of the entire GNPS revealed an order-of-magnitude jump in the number of PNP variants compared to all previous PNP discovery efforts. Moreover, VarQuest revealed a surprising diversity of PNPs that may reflect evolutionary adaptation of various bacterial species to changing environment and competition, e.g., a continuous change in the repertoire of variants of peptidic antibiotics in response to developing antibiotic resistance.

“Researchers in the field of natural products are collecting large-scale metabolomic data from microbial strains,” said paper co-author Hosein Mohimani, Pevzner’s former student who is now an assistant professor in the Computational Biology Department at Carnegie Mellon University. “Natural product discovery is turning into a highly data-intensive field, and the area has to get prepared for this transformation in terms of making sense of Big Data. VarQuest is the first step toward making sense of the Big Data already collected in the field.”
Prior to joining the Carnegie Mellon faculty in October 2017, Mohimani was a project scientist in the Computer Science and Engineering department at UC San Diego, working under Pevzner in the NIH-funded Center for Computational Mass Spectrometry. Mohimain is also a UC San Diego alumnus (Ph.D. ’13) from the university’s Electrical and Computer Engineering department.
__________
*Alexey Gurevich, Alla Mikheenko, Alexander Shlemov, Anton Korobeynikov, Hosein Mohimani & Pavel A. Pevzner, Increased diversity of peptidic natural products revealed by modification-tolerant database search of mass spectra , Nature Microbiology (2018) doi:10.1038/s41564-017-0094-2
Media Contacts
Doug Ramsey, (858) 822-5825, dramsey@ucsd.edu
Related Links